Table of Contents
Overview of ADCI_Online
ADCI_Online is a subscription service which grants access to ADCI on a dedicated cloud-based system through a web browser. Interacting with ADCI “on the cloud”:
- Eliminates the need for a dedicated computer system to run ADCI
- Generates the same results as dedicated systems
- Has a short-term subscription format which reduces cost
- Can accommodate on-demand bursts of computing power when necessary
- Can ensure that cytogenetic data is colocated in the same region as the user
- Securely isolates individual user data and protects software analysis from intrusion, disruption and corruption
- Allows dose estimation to be carried out anywhere there is a reliable internet connection
Scalable
Although each system running ADCI_Online has less computing power than a high-performance system running ADCI Desktop, the cloud-based nature of ADCI_Online allows for rapid expansion of resources.
If many samples must be processed quickly, more computing power can be added (incurs additional cost). CytoGnomix can quickly reconfigure ADCI_Online within ~15 minutes to:
- Clone the system
Each ADCI_Online streaming instance is cloned from a single base image. This image can be cloned as many times as necessary, resulting in an array of cloud-based systems available for use. - Increase the computing power of each streaming instance
The base image can be loaded on a more powerful system when it is cloned at the start of a session. This means the streaming instances will be running on more powerful hardware. - Both of the above
A similar array of systems could be achieved by utilizing multiple physical computers, however the ability to expand and reduce available resources as necessary provides flexibility and reduces costs.
Data Security
An Amazon Web Services (AWS) S3 Bucket is used to store metaphase image files and reports generated by analysis of the data. A user-specific directory is created within the S3 bucket and is named according to the SHA-256 hash of the their e-mail address, thus e-mail addresses are not visible to other users. The user-specific directory in the S3 Bucket is mounted to the cloud-based system on which ADCI is executed, granting the user access to their uploaded images in ADCI_Online.
Files are encrypted in transit to/from the Bucket (HTTPS protocol) and server-side encryption is applied to all files in the Bucket (AWS Key Management Service). File types (e.g. images) are verified by the system upon uploading. Uploaded files are prevented from running as executables on AWS AppStream. Internal elements of ADCI software are invisible/inaccessible to the user. Internet and browser access from within ADCI_Online itself is disabled. Only files created by ADCI_Online can be downloaded by the UserName that generated them.
Uploaded metaphase cell image data are owned by and may be removed by each respective user. Automated program scripts delete user data 1 month after uploading, unless the user makes arrangements with CytoGnomix to maintain it. CytoGnomix will not work with it or keep it elsewhere, unless the user requests that the company provide consulting services.
User Experience
Before a subscription period begins, a new user can sign into our Javascript web app (sign-in credentials provided) and upload metaphase images to cloud storage. This mechanism is also used to download ADCI reports after they have been generated. Consult the cloud storage wiki page for a more detailed description of this process.
To access ADCI_Online, the user signs into AppStream in their web browser (sign-in credentials provided) and requests a new streaming session. Behind the scenes, a new streaming instance is cloned from a snapshot of the ADCI_Online system. Simultaneously, the user’s S3 storage directory is mounted to the streaming instance, allowing the user to access their uploaded metaphase images and save results generated while executing ADCI. Consult the accessing ADCI_Online wiki page for a more detailed description of this process.
Remote Execution of ADCI
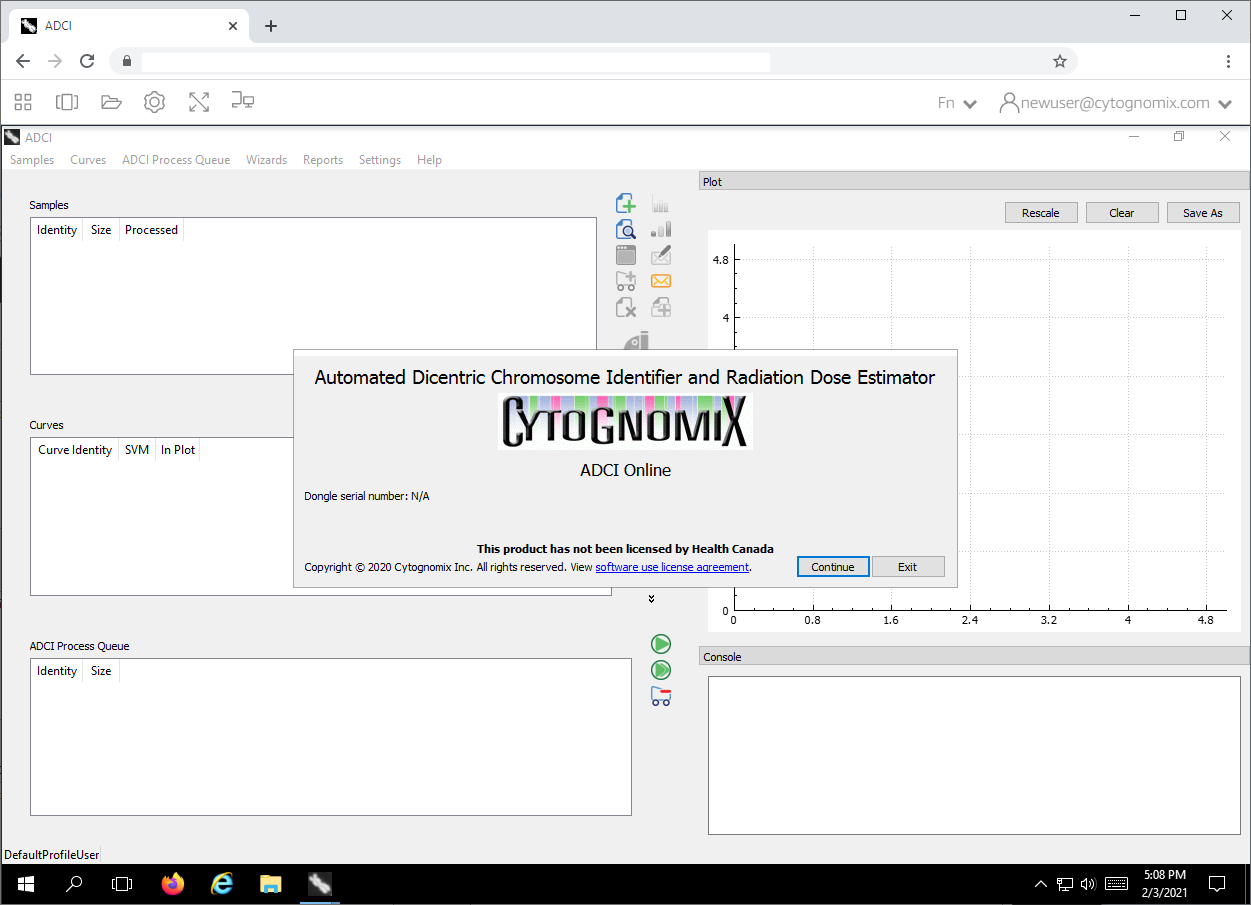
ADCI_Online runs exclusively on the cloud-based system and pixels are streamed to the user through a web browser (shown in image to the right). Therefore, computing resources and disk space are not consumed on the local system. Local keyboard and mouse commands are sent to the cloud-based system to control ADCI_Online.
Operationally, ADCI_Online is indistinguishable from the MS Windows version that runs on a dedicated, standalone computer. Because the base hardware configuration of ADCI_Online is a 2 processor CPU, 3.75Gb RAM, sample processing is ~3 fold slower than the recommended Desktop computer running ADCI (Intel i7-7th gen 4 proc. CPU, 16Gb RAM). However, as mentioned, ADCI_Online can be rapidly scaled to meet “burst” requirements such as an emergency situation requiring examination of many samples.